Scientists at the Berlin Institute of Health (BIH) and Charité – Universitätsmedizin Berlin, working with colleagues from the United States, have selectively modified the control regions of 20 disease-relevant genes. This enabled them to identify precisely those modifications that have the greatest influence on disease state. Their findings will now allow physicians to predict which DNA modifications found in patients are actually responsible for disease, thus highlighting them for a potential targeted therapy. The researchers have just published their results in the journal Nature Communications.
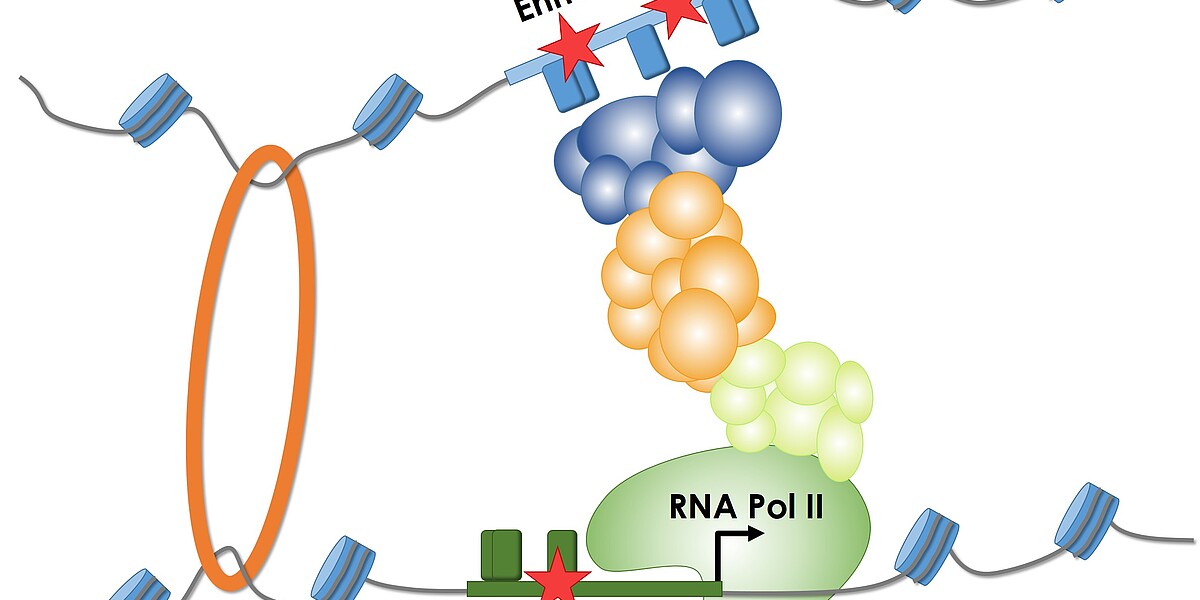
Many diseases occur because a person’s DNA contains errors, so-called mutations. These cause vital protein molecules to be produced incorrectly. Sometimes the molecules are altered to such an extent that they are no longer able to perform their function. But more often, modifications in the gene control regions (regulatory sequences, also known as promoters and enhancers) cause the wrong amount of protein to be produced. The cells produce either too much or too little protein, or none at all. “That can, for example, result in cancer if a type of protein that promotes cell division is produced in too great a quantity,” explains Martin Kircher, head of the BIH Junior Research Group on Computational Genome Biology and lead author of the paper.
Yet cancer cells are precisely where many mutations often occur – and some of these mutations have no impact while others actually cause or speed up the disease by influencing how much protein is produced. Before physicians prescribe a therapy that specifically combats certain mutations, they should know what role these play for the disease.
To help with this, a team of scientists around Dr Kircher took the control regions of 20 disease-relevant genes and modified them piece by piece – or to be more precise, the DNA base by base. For this purpose they developed a method that was able to perform these modifications in a high-throughput manner and allowed them to test them simultaneously. Using cell cultures, they examined what impact each modification had on protein production. “Some 85 percent of modifications have no measurable effect, while two-thirds of the remaining 15 percent reduce the amount of protein produced,” reports Dr Kircher. And the extent to which they effect cell activity is heavily dependent on the individual mutation and the control region investigated: “Replacing a base with another one usually has less impact than completely removing the base.”
The control elements investigated were from genes that are modified in patients suffering from cancer, heart failure, genetically high cholesterol, or various rare diseases. The researchers have made the results of their mutation analyses – more than 30,000 in total – freely available to anyone on the internet. Martin Kircher now hopes that this treasure trove of data will be used: “It would be great if physicians who have analyzed their patients’ DNA would consult our database to find out what effect the identified mutation, in all likelihood, has. This will enable them to assess whether the modification found in patients is suitable for a targeted therapy.”
The considerable time and effort needed to test 20 of possibly hundreds of thousands of control regions raises the question of whether machine learning or artificial intelligence could help predict which mutations produce which effects. There are already various computer programs that are trying to do exactly that. Dr Kircher and his colleagues therefore also studied how well different programs could predict the modifications observed in the cell culture. “The results were unfortunately very disappointing,” reports the Bioinformatician, “because the predictions rarely corresponded with our observations. Sometimes they even predicted the exact opposite.” The scientists now hope that their comprehensive data set can also be used to improve such predictive software.